Chapter 11 Data Classes & Methods
Construction of complex objects
So far we have a series of different data structures/objects including vectors, matrices, lists and data frames. We have learned that ‘more complex objects’ (matrices; data frames) are all based on the most basic objects (vectors; lists). This idea can be used to create a variety of additional (even custom) data classes for specific purposes.
Generic functions
We observed that the behaviour of a function may depend on a class of an object we call it for. This happens for generic functions, such as:
print(),str(): presenting an object in a way which is meaningful for its content.summary(): showing a summary of characteristics specific to an object.plot(): creating a plot suitable for particular objects (single vector, two vectors, formula, data frames).
Besides these four examples, a range of additional generic functions exist. If we need something very specific for our own work we can also create custom classes and methods for specific data types or define additional generic functions (if needed). This is relatively easy once we understand how it works in R.
Goal of this chapter
In this chapter, we will learn about two additional data structures in R.
One for categorical data (factor) and date-time objects (Date, POSIXt).
First, factors are introduced and ‘deconstructed’ to see how an object
of class factor is constructed. This helps to understand how generic
functions work and allows us to write our first own custom class.
To back this up we will learn how date and date-time objects in R work
and how they are constructed.
11.1 Categorical data (factors)
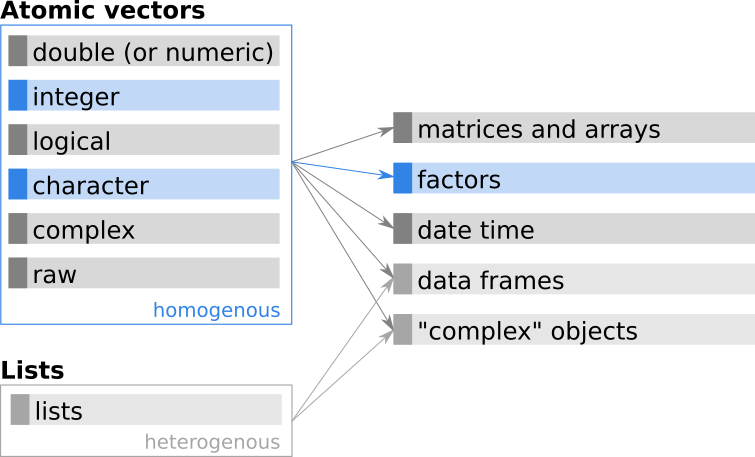
Figure 11.1: Additional base R classes based on atomic vectors.
Categorical data are used for qualitative variables when the outcome of a specific observation/measurement falls into (exactly) one of a number of known and countable categories.
Clarify the jargon: In R, objects for categorical data are called factors, while the different categories in which an observation can fall is called a level. Other programming languages/software may use different labels, e.g., ‘Enumerated Type’ (ENUM) in SQL/MySQL databases, C/C++, and JAVA, or ‘category’ in Python pandas.
Binary examples: Only two distinct outcomes (two possible categories).
| Description | Levels | |
|---|---|---|
| Nominal | Precipitation | "rain", "no rain" |
| Power supply | "on", "off" |
|
| Purchase of a product | "yes", "no" |
Examples with \(\ge 3\) categories: When having more than two possible outcomes, one talks about categorical data (whereof binary is a special case). Categorical data can be nominal (unordered) or ordinal (ordered). Ordered means that we can bring the categories in a distinct order, which can not be done for nominal data.
| Description | Levels | |
|---|---|---|
| Nominal | Gender | "female", "male", "diverse" |
| Blood type | "A", "B", "AB", "0", … |
|
| Means of transport | "bike", "car", "train", … |
|
| Ordinal | Education | "bachelor", "master", "phd" |
| Satisfaction | "low", "medium", "high" |
|
| Rating | "AAA", "AA+", "AA", … |
Nominal versus ordinal:
- Gender (nominal): We cannot bring ‘gender’ into a specific order, males are not ‘below/above’ females or diverse persons.
- Education (ordinal): The three categories ‘bachelor’, ‘master’, and ‘phd’ can be brought in a distinct order. We cannot get a Masters degree as long as we have not finished our Bachelors study. And we need a Masters degree first to finish our PhD (doctorate).
Creating factors
Factors can be created using the function factor() (the constructor function).
From the documentation:
Important arguments:
x: a vector of data, usually taking a small number of distinct values.levels: an optional vector of the unique values (as character strings) that ‘x’ might have taken. The default is the unique set of values taken by ‘as.character(x)’, sorted into increasing order of ‘x’.labels: either an optional character vector of labels for the levels (in the same order as ‘levels’ after removing those in ‘exclude’), or a character string of length 1. Duplicated values inlabelscan be used to map different values ofxto the same factor level.exclude: a vector of values to be excluded when forming the set of levels. This may be a factor with the same level set asxor should be acharacter.ordered: logical flag to determine if the levels should be regarded as ordered (in the order given).
From character vector
When a factor is created from a character vector, all unique values of the vector are taken as levels (categories), the categories are lexicographically sorted.
Let us create a character string which contains the educational attainment of a series of people, coded by the highest university degree.
## [1] "master" "master" "bachelor" "phd" "bachelor" "master"## [1] "character"This is a classical categorical variable: We have 6 observations (or elements), but only 3 possible levels (categories). Thus, let us convert this character string into an object of class factor.
## [1] master master bachelor phd bachelor master
## Levels: bachelor master phd## [1] "factor"The object degree is now of class factor and the representation
of the object changed – not only the quotation marks are gone (""s), we
also get an additional line with Levels. Each outcome of our original
character vector is taken as one of the levels, lexicographically sorted
(try sort(unique(degree)) on the character vector).
Plotting factors & numerical summary:
Before going into more detail let us see why factors are useful.
Let us have a look at the numerical summary (summary()).
## bachelor master phd
## 2 3 1As R recognised that we have categorical variables we now get absolute
counts (as from table()). The same if we use the generic function plot().
As a line plot would not be meaningful, R automatically draws a bar plot.
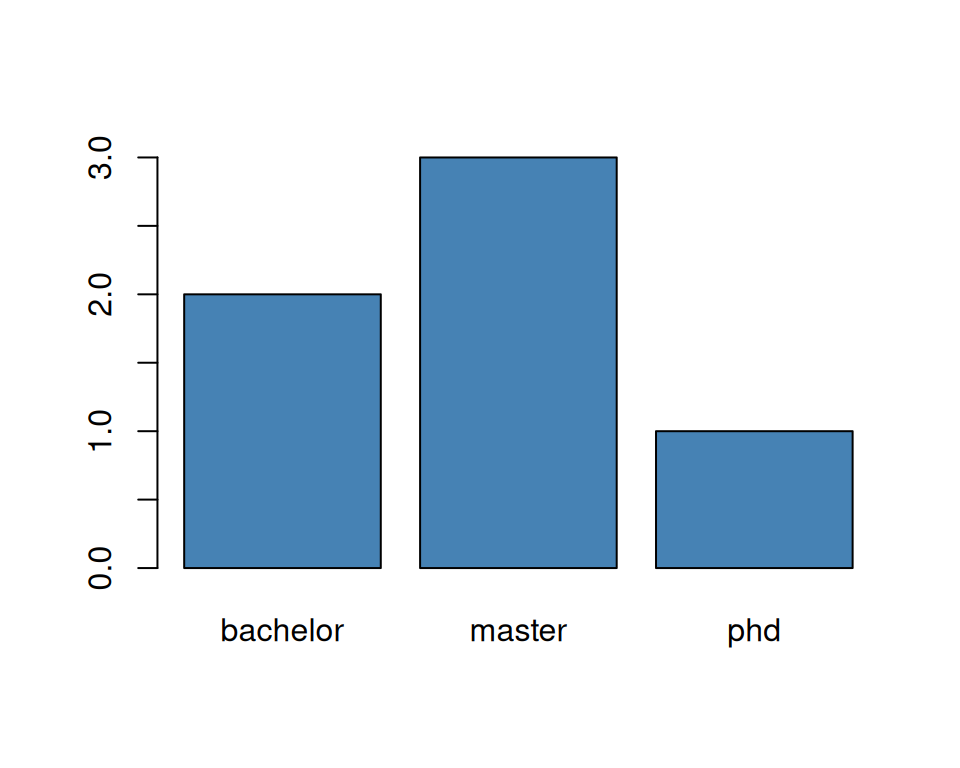
As you can see, factors help us a lot when dealing with categorical data.
Basic properties and attributes:
- Properties: Like all objects, factor objects always have a length, and a type.
- Attributes: Factor variables always have two attributes, a class (
factor) and an attribute levels.
Let us investigate the object degree from above.
## $class
## [1] "factor"
##
## $typeof
## [1] "integer"
##
## $attributes
## $attributes$levels
## [1] "bachelor" "master" "phd"
##
## $attributes$class
## [1] "factor"While the class is as expected, typeof() returns us integer. Keep
this in mind (will become clear later). In addition, we can see
that we have the two (mandatory) attributes "class" (simply "factor") and "levels",
the character labels (names) of the different levels/categories.
Dedicated extractor functions: For factor variables, a series of extractor functions exist we have not seen so far.
levels(): Extracts the character labels (levels); returns a character vector.nlevels(): Returns the number of different levels; single integer.
## $levels
## [1] "bachelor" "master" "phd"
##
## $nlevels
## [1] 3Extracting attributes: In the meantime we learned how to work with lists.
And we know that all attributes of R objects are stored as a named list
(check the output of attributes()).
We can access attributes by using attr(x, which) where x is the target
object, and which the name of the attribute, or by using attributes(x)
(returns named list). Thus, we could also use the following commands to get the
same information as levels() and nlevels() using:
## [1] "bachelor" "master" "phd"## [1] "bachelor" "master" "phd"## [1] 3## [1] 3Note: It’s recommended to use the dedicated extractor functions
(here levels() and nlevels()) if available.
Structure of a factor: Let us have a closer look to the structure of a factor object and how it is constructed.
## Factor w/ 3 levels "bachelor","master",..: 2 2 1 3 1 2As shown, we get the information that we have a factor variable with (w/) 3 levels; followed by names of the first few levels. What about the integer values at the end of the output?
Let us use a new function called unclass(). It removes the class of an object
and deconstructs it to the simplest possible object, a vector (atomic vector) or list
(generic vector).
## [1] 2 2 1 3 1 2
## attr(,"levels")
## [1] "bachelor" "master" "phd"We can now see how the object is constructed. A factor object is nothing else
than an integer vector ([1] 2 2 1 3 1 2)
with an additional attribute levels containing the names of the three levels.
This is nothing else as a ‘lookup table’. The integer vector (index vector) tells
us that the first entry of our object belongs to category 2,
the second to category 2 and so far and so on.
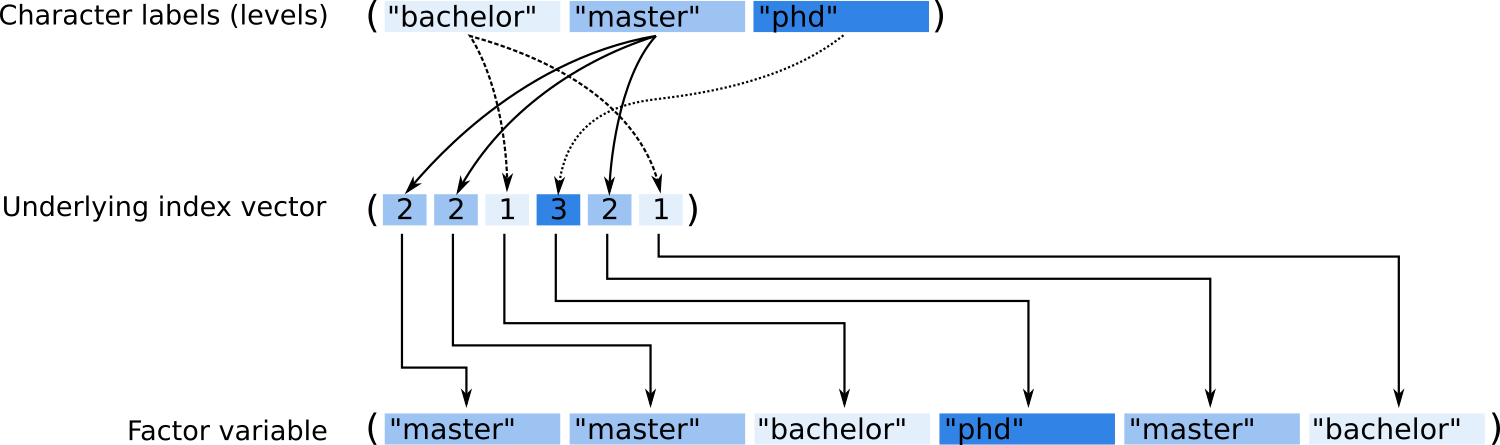
Figure 11.2: Graphical representation of the internal structure of a factor variable.
As our factor variable is indexed by index values (our integer vector), we can also use this for subsetting. An example:
## [1] "MSc" "MSc" "BSc" "PhD" "BSc" "MSc"… this is nothing else than subsetting by index (see Vectors: Subsetting by index).
By hand: Let us do c("BSc", "MSc", "PhD")[degree] ‘by hand’ to better
understand what’s going on.
- Crate yourself a new object
new_names <- c("BSc", "MSc", "PhD")(character vector) - Coerce (convert)
degreeto integer using explicit coercion, store result onidx. - Use
idxandnew_namesto create the same as shown above.
Solution. Let us create the new character vector and the idx object in the first code chunk.
The coercion is done using as.integer(degree).
## [1] 2 2 1 3 1 2We can now use idx to subset by index as follows:
## [1] "MSc" "MSc" "BSc" "PhD" "BSc" "MSc"… and that’s exactly what is shown above:
## [1] "MSc" "MSc" "BSc" "PhD" "BSc" "MSc"Summary:
- The number of categories (levels) is fixed.
- Character labels (names) of the levels/categories are stored as attribute
"levels". - The data are indexed by an integer vector (a factor object is an integer vector with an attribute levels).
- The separate class (
factor) allows the creation of dedicated methods for categorical data such as subsetting, visualization, modelling, …
From integer vector
Instead of starting from a character vector, we can also create factor variables based on an integer vector. This is often useful when we have a data set at hand which contains categorical information coded as integers as in the following example.
## [1] 1 3 1# Convert to factor variable
(degree2 <- factor(degree2,
levels = 1:3,
labels = c("bachelor", "master", "phd")))## [1] bachelor phd bachelor
## Levels: bachelor master phdAs shown in the comments, we know that this data set may contain
three possible categories. Level 1 corresponds to "bachelor", 2 to "master",
and 3 to "phd". The two arguments levels and labels define this connection
and are used to convert our integer vector into a proper factor variable.
Note: We have defined three possible outcomes, even if the data set itself only
contains two of them (we have no 2 = master).
Creating ordered factors
So far we always had nominal data (unordered). If the order of the different
categories is important, we can set the additional argument ordered to TRUE.
The rest is the very same as above, however, the output does look different:
# Calling the object 'odegree' (ordered degree)
(odegree <- factor(c(1L, 3L, 1L),
levels = 1:3,
labels = c("bachelor", "master", "phd"),
ordered = TRUE))## [1] bachelor phd bachelor
## Levels: bachelor < master < phdAs shown, we now get Levels: bachelor < master < phd;
The ‘smaller then’ operator (<) indicates that the categories are now
ordered (from left to right). This adds some additional functionality:
## [1] TRUE## [1] FALSE… which will not work on nominal (unordered) factors. In addition, such ordered factors can be very useful when estimating statistical models (not part of this course).
Factors from continuous numeric
Another very handy function is cut(). It allows us to
create a categorical variable (factor) based on defined intervals based
on a continuous scale (numeric vector).
Usage:
cut(x, breaks, labels = NULL,
include.lowest = FALSE, right = TRUE, dig.lab = 3,
ordered_result = FALSE, ...)Important arguments:
x: a numeric vector which is to be converted to a factor by cutting.breaks: either a numeric vector of two or more unique cut points or a single number (greater than or equal to 2) giving the number of intervals into whichxis to be cut.labels: labels for the levels of the resulting category. IfFALSEinteger codes are returned (instead of factor).ordered_result: logical: should the result be an ordered factor?
Example: We would like to classify the state of water given a specific
temperature in degrees Celsius. Below \(0\) degrees Celsius "solid", \(0 > x \le 100\)
degrees Celsius as "liquid", and \(> 100\) degrees Celsius "gas".
We can specify the breaks (where to cut the continuous variable) using the
breaks argument. If set to a single integer, R selects some breaks (e.g., 8 breaks).
In case we know where to cut, as in this example, we can define a numeric vector
(on breaks) defining these break points.
temp <- c(17.5, -4.3, 36.0, 68.1, 130.0) # observations
cut(temp,
breaks = c(-Inf, 0, 100, Inf), # 4 breakpoints = 3 segments
labels = c("solid", "liquid", "gas")) # 3 labels (categories)## [1] liquid solid liquid liquid gas
## Levels: solid liquid gasTake care: if we have 4 breaks we define 3 segments (categories) and
therefore only need 3 labels. If no labels are specified, R will set
default levels of the form (a, b] (mathematical definition of an interval).
The excursion below (not mandatory) shows more details on that for those interested.
By default, the levels are set to (a, b] which
mathematically defines a ‘left open, right closed’ interval. This simply means
that this interval includes all numbers \(a < x \le b\) (\(a\) not included; \(b\)
included). In our example, an observation falling on 0 would fall into the
first segment, those on 100 into the second segment.
This can be changed by setting right = TRUE creating ‘left closed, right open’ intervals.
## [1] (0,100] (-Inf,0] (0,100] (0,100] (100, Inf]
## Levels: (-Inf,0] (0,100] (100, Inf]In addition: the lowest (or largest if right = TRUE) value will be ignored
except you set include.lowest = TRUE. This can be crucial as well but is no
problem in this example, as the outer bounds have been set to -Inf, and
+Inf.
# Using absolute 0 (0 Kelvin) as the lower bound,
# Include lowest (in case it would be -273.15).
(state <- cut(temp, # numeric values
breaks = c(-273.15, 0, 100, Inf), # 4 breakpoints
labels = c("solid", "liquid", "gas"), # 3 labels (categories)
include.lowest = TRUE, # include lowest (-273.15 as 'solid')
ordered = TRUE)) # ordered categories## [1] liquid solid liquid liquid gas
## Levels: solid < liquid < gaslist(class = state,
levels = levels(state),
nlevels = nlevels(state),
is.ordered = is.ordered(state))## $class
## [1] liquid solid liquid liquid gas
## Levels: solid < liquid < gas
##
## $levels
## [1] "solid" "liquid" "gas"
##
## $nlevels
## [1] 3
##
## $is.ordered
## [1] TRUE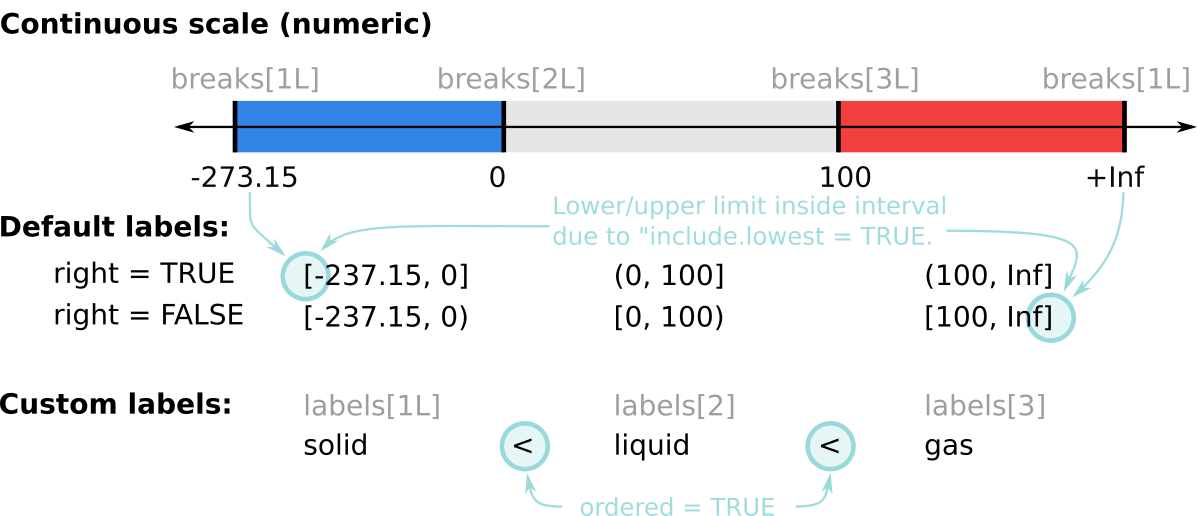
Figure 11.3: Graphical representation of the function cut(); three intervals (-273.15 to 0, 0 to 100, and 100 to +Inf).
Subsetting and replacing elements
Factor variables allow for proper subsetting or to replace specific elements, the same as for vectors. This includes subsetting by index, subsetting by logical vectors, and even subsetting by name (it is technically possible to add names to the elements of a factor; not seen often).
Subsetting by index: Extract the first element, and elements c(6, 3) (in this order).
The result is still a factor, all levels are kept.
## [1] master
## Levels: bachelor master phd## [1] master bachelor
## Levels: bachelor master phdSubsetting by logical vectors: Straightforward.
## [1] master master
## Levels: bachelor master phdReplace elements: We can also assign levels and overwrite specific elements. Note: Only works for defined levels!
degree[5] <- "bachelor" # replaces element 5
degree[6] <- "habilitation" # won't work, not a valid level!## Warning in `[<-.factor`(`*tmp*`, 6, value = "habilitation"): invalid factor
## level, NA generatedSubsetting and replacing levels
In the same way, we can also subset or replace (specific) levels and change the names of the categories if needed.
## [1] "bachelor"## [1] MSc MSc BSc PhD BSc <NA>
## Levels: BSc MSc PhDReference category, unused levels
Sometimes it is necessary to have a very specific category as the first level in your factor object, e.g., when estimating statistical models. This first level is called the reference category or reference class. Furthermore, it might sometimes be useful to remove (drop) unused levels. The following excursion (not mandatory) shows how this can be done for those who want to know.
We know that, by default, the levels of a factor are lexicographically sorted (if not ordered). Imagine you perform a medical experiment testing some drugs. The participants either get the “standard” treatment, the “new” treatment, or a “placebo” medication (with no effect).
## [1] placebo new placebo standard new
## Levels: new placebo standardOur first level is new, but we want placebo as the first level
as this is our reference category (ref; first category/level). This is often
important when using such data for statistical modelling (beyond this course).
To define placebo as our first category, we can use relevel().
## [1] placebo new placebo standard new
## Levels: placebo new standardDrop unused levels
In case you have unused levels and you want to drop them, call droplevels().
Let us take the example from above with c("bachelor", "master", "phd") where
we have not had any master student (see From integer vector):
## [1] bachelor phd bachelor
## Levels: bachelor master phdLet us drop all unused levels: the result is a factor object with only two
levels ("bachelor", and "phd").
## [1] bachelor phd bachelor
## Levels: bachelor phdNote: For unordered factors (orderered = FALSE;, default)
droplevels(degree2) does the very same as factor(degree2).
droplevels(degree2) is a convenience function which, internally, calls factor().
What factor(degree2) does: it converts the input (degree2) into a
character vector (explicit coercion), and creates a new factor object out of it.
If we have a level but no observation assigned to this level/category, it will not be included in the character vector. Thus, when creating a new factor out of it, it will be dropped. By hand:
## [1] "bachelor" "phd" "bachelor"## [1] bachelor phd bachelor
## Levels: bachelor phd## [1] bachelor phd bachelor
## Levels: bachelor phd## [1] bachelor phd bachelor
## Levels: bachelor phdFor ordered factors this will not work as as.character() converts
our data into a bare character vector – wherefore we will again
end up with lexicographically ordered levels.
Methods for factors
Question: We have now seen what we can do with factors. But is this really necessary? I mean, couldn’t we simply take an integer or character vector and have the very same information?
Answer: Yes, but there are many advantages of having a dedicated class for categorical data and R provides a series of methods for data handling (e.g., coercion, subsetting, visualization, …).
Let us highlight the differences between a pure integer vector, a character vector,
and a factor object (containing the ‘same’ information).
To do so, we first coerce our factor degree into a pure integer vector (degree_int)
and a character vector (degree_chr).
degree <- factor(c("master", "master", "bachelor", "phd", "master", "bachelor"))
degree_int <- as.integer(degree) # coerce to integer
degree_chr <- as.character(degree) # coerce to characterWe will then use the generic functions print(), summary() and plot()
to highlight the advantages: The good, the bad, and the ugly.
The good
print(): Easy to read as all values are coded with their corresponding category. In addition, we see what categories we have.summary(): As we have counts, the numerical summary shows us how many observations we have in each class (absolute counts; astable()).plot(): As seen above we get a barplot as a line plot or scatter plot would not make any sense.
## [1] master master bachelor phd master bachelor
## Levels: bachelor master phd## bachelor master phd
## 2 3 1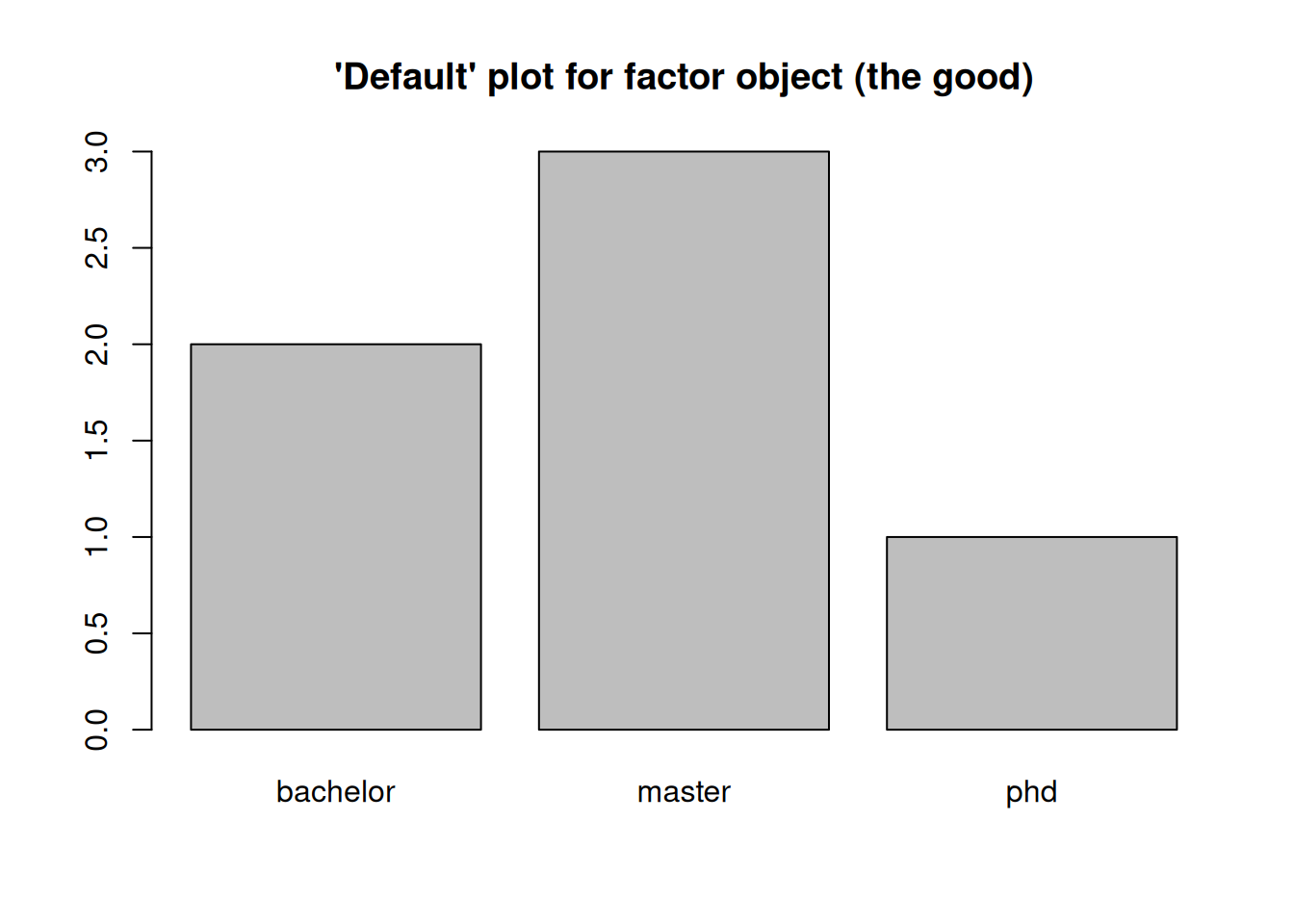
The bad
When using the integer sequence, R handles the object as a numeric vector. However, standard arithmetic on categorical data is not meaningful! The arithmetic mean or median (as well as minimum/maximum and quartiles) cannot be interpreted!
print(): Not easy to interpret, we would need a lookup-table to decode the information.summary(): Standard arithmetic is not very meaningful for categorical data.plot(): Not meaningful, especially not for larger objects.
## [1] 2 2 1 3 2 1## Min. 1st Qu. Median Mean 3rd Qu. Max.
## 1.000 1.250 2.000 1.833 2.000 3.000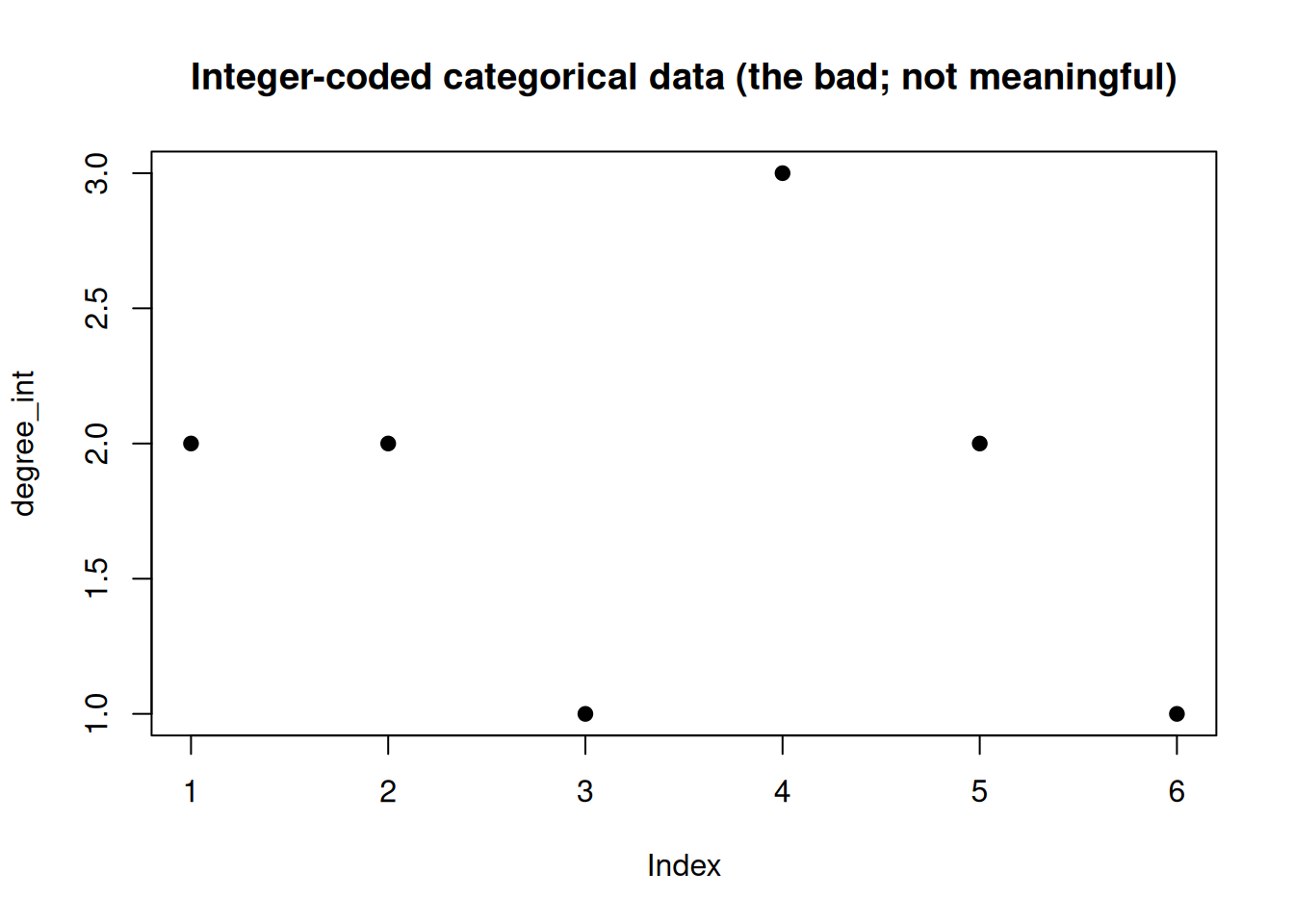
The ugly
print(): Easy to read, except that we don’t see which categories exist. Otherwise, similar to the output of the factor.summary(): Does not give us meaningful information.plot(): Would throw an error, we cannot plot a character vector.
## [1] "master" "master" "bachelor" "phd" "master" "bachelor"## Length Class Mode
## 6 character characterComputational advantage
In addition, a factor variable has one more advantage. An integer vector of length \(N\) requires much less space on your hard drive and in memory, and calculations on integer vectors can be much faster than on character vectors. However, plain integer vectors are less informative (for humans) as readable labels.
Best of both worlds: Factor variables combine the advantages of both; The data are stored as an integer (efficient), the labels have only be stored once as character; while we, as users, still have the advantage of human-readable labels/categories.
11.2 Object orientation
This strategy that we have discussed above for factors can also be applied more generally:
- Definition of a suitable class for objects that are then required to have a certain structure.
- Providing methods to generic functions for capturing typical computations or workflows based on these objects.
- Possibility of organizing classes in a hierarchy so that a child class inherits the behavior from the parent class.
This strategy is sometimes also referred to as object orientation or object-oriented programming (OOP) in the R ecosystem. However, a caveat is that the way this object orientation is implemented in base R is rather different and much more informal compared to other object-oriented languages like C++, Java, or Python. It is also rather different from OOP definitions and discussions in standard computer science textbooks.
This is also the reason that R in fact comes with several object orientation systems, including some that are closer to more standard OOP implementations:
- S3: This is the first and simplest object orientation system in R. It is based on generic functions for classes like the factors introduced above.
- S4: A somewhat more formal approach, also based on generic functions, but with stricter validity checks of the objects.
- Reference classes (RC): More formal OOP implementation where methods belong to the classes/objects directly (and not to generic functions).
- R6: An OOP implementation similar to RC but intended to be more lightweight.
Subsequently, we confine ourselves to the basic S3 system which is used in R’s basic statistics functionality and in most commonly-used packages on CRAN. It is elegant in its minimalism and hence relatively easy to adopt for the creation of new classes and methods. Hence subsequently only S3 is introduced in some more detail before showing how custom classes and methods can be created with it.
S3 generics
S3 is based on so-called generic functions. A generic function is a regular
function (see Chapter Functions) with a
series of arguments and a call to UseMethod() with the name of the generic function
in the instruction-part of the function.
This tells R that there are specific methods for specific objects, depending on
the class of the object. Let us have a look to the insides of the generic
function summary by calling summary (without brackets; shows the function definition).
## function (object, ...)
## UseMethod("summary")
## <bytecode: 0x55a93250b390>
## <environment: namespace:base>If we ignore the last two lines we basically have the following function:
We have seen several times what the summary() function returns, and
that the returned data summary depends on the class of the object
(numeric vectors, character vectors, matrices, …) given as the first input argument.
The function above is not doing that at all. Instead, this is the generic function.
UseMethod("summary") tells R that there must be other functions – now
called methods – for objects of different classes.
Arguments
object(required): The main argument when callingsummary(). The class of this object determines the dispatch to methods....: An arbitrary number of optional arguments passed on to the methods.- Any method must implement these two arguments (
object,...) but may contain further arguments.
Details
- When
summary()is applied to an object of class"foo", R tries to call the functionsummary.foo(). If it does not exists, it will callsummary.default(). - R objects can also have a vector of classes, e.g.,
c("matrix", "array"), meaning that the object is of class"matrix"inheriting from"array"("matrix"is a special version of the more generic"array"). - In this case, R
first tries to apply
summary.matrix(), then (if not existing)summary.array(), and then (if both do not exist)summary.default().
Using methods
- Methods defined for a certain generic function can be queried using
methods(). methods(summary)returns a (long) list of all methods available includingsummary.factor()andsummary.default()(but notsummary.numeric(), assummary.default()would be applied to objects of classnumeric).- As it is not recommended to call methods directly, some methods are marked as being non-visible to the user and these cannot (easily) be called directly.
- Even if visible, it is preferred to call the generic, i.e.,
summary(g)instead ofsummary.factor(g)!
Typical generic functions
print(),plot(),summary(),str(), which print, plot, summarize, and describe the structure of an object are available for many classes.- For data classes often:
[,[<-,c()(for subsetting and combining objects), coercion functionsas.numeric(),as.character()etc. Possibly alsoformat(), mathematical operations etc. - For statistical models (not part of this course; e.g., from
lm()):predict(),coef(),residuals(), and many more to do predictions, extract estimated coefficients, or get the residuals of the model.
See what’s defined We can use methods(generic.function, class) to get
the available methods for a certain generic function (as above), or a specific class.
Let us see which methods exist for droplevels(). The output below tells us
that there is a method for data frames (droplevels.data.frame) and one for
factors (droplevel.factor) as shown previously.
## [1] droplevels.data.frame droplevels.factor
## see '?methods' for accessing help and source codeWe might also be interested in what methods exist for an object of a specific class,
here class "factor":
## [1] Arith Compare Logic Math Ops
## [6] Summary [ [<- [[ [[<-
## [11] all.equal as.Date as.POSIXlt as.character as.data.frame
## [16] as.list as.logical as.vector c cbind2
## [21] coerce droplevels format initialize is.na<-
## [26] kronecker length<- levels<- plot print
## [31] rbind2 relevel relist rep show
## [36] slotsFromS3 summary xtfrm
## see '?methods' for accessing help and source codeBesides plot(), summary(), and droplevel() we can also see that there
is a generic function relevel() which we already used earlier.
Source code: We can always look into the definition of the S3 methods
using getS3method(f, class) (f: function name, class: class name).
## function (x, exclude = if (anyNA(levels(x))) NULL else NA, ...)
## factor(x, exclude = exclude)
## <bytecode: 0x55a93a0161a8>
## <environment: namespace:base>If exported (not all methods are exported) we can get the same result by
typing droplevels.factor (no brackets at the end).
Note: Not all methods are implemented for every possible object.
Nor are all functions generic. Just as an example, there is no method
to combine factor variables (c(a, b) where a and b are factors).
Which does not mean we can not implement it ourself! The exercise
below tries to demonstrate that.
Combining factors If we simply try c() we will end up with a messed up
result as R only combines the two underlying integer vectors.
degree <- factor(c("master", "bachelor", "bachelor"))
degree2 <- factor(c("phd", "bachelor", "bachelor", "phd"))
c(degree, degree2) # Losing information!!## [1] master bachelor bachelor phd bachelor bachelor phd
## Levels: bachelor master phdWe could, however, do this manually by:
- Coercing both factor objects (
degreeanddegree2) to character. - Combine the two character vectors.
- Create a new factor out of it.
This would look as follows:
## [1] master bachelor bachelor phd bachelor bachelor phd
## Levels: bachelor master phdExercise: Instead, we could write a method for that. c() is a generic
function, thus we can write a custom c.factor() method ourselves.
- Write a function
c.factor()(method) with two input arguments. - Internally, create a new combined factor object.
- Return the new object.
- Test
c(degree, degree2)with the two objects above.
Solution. All we need to do is the following:
c.factor <- function(x, y) {
return(factor(c(as.character(x), as.character(y))))
}
c(degree, degree2)## [1] master bachelor bachelor phd bachelor bachelor phd
## Levels: bachelor master phdEt voila, we have a dedicated method to combine factor objects. c(degree, degree2) returns an object of class factor combining both input arguments.
And yes, the function above might need some improvements (just for
illustration). This is the basic idea of S3 and makes our life easier.
Generalization: The simple demo method (c.factor) above
only works for \(2\) inputs. We could generalize the function
such that it works with \(\ne 2\) arguments.
Note that this is way beyond what you need to know, but we can
write our c.factor more generic. This uses a series of functions
you have not seen (and you don’t have to know)!
# Takes '...' (a list of 1, 2, or N objects),
# converts the content to character and creates
# a new combined factor object.
c.factor <- function(...) {
return(factor(unlist(lapply(list(...), as.character))))
}
c(degree, degree2) # would also work with 3 or more arguments## [1] master bachelor bachelor phd bachelor bachelor phd
## Levels: bachelor master phd11.3 Creating a custom class
So far we have seen how existing classes work – but we can also create our own ones! As motivation: We will use this to handle the air pollution (Luftverschmutzung) data set of Beijing, China. Some big Chinese cities are known for very bad air pollution which can cause serious health issues. We would like to write a class which tells us how often a specific limit (or threshold) was exceeded.
Goal: We will implement a new class called threshold for handling numeric
values in combination with a specific limit. Our object should have the
following properties, attributes, and methods:
- Object properties:
- Type:
double(numeric values). - Length: any length (vectors of length \(\ge 1\)).
- Type:
- Object attributes:
- Class:
threshold. - limit: single numeric value; our predefined limit.
- Class:
- Methods:
as.character()/print(): Customas.characterandprintmethods for now.
As there is no formal definition for S3, all we have to do is to assign
a new class to the object pm25, and the threshold we want to use.
pm25 <- c(100, 75, 230, 220, 50) # Vector for testing
attr(pm25, "limit") <- 150 # Adding attribute 'limit'
class(pm25) <- "threshold" # Adding class attribute
print(pm25)## [1] 100 75 230 220 50
## attr(,"limit")
## [1] 150
## attr(,"class")
## [1] "threshold"We can now write our custom methods with the naming convention
genericFunction.className(), e.g., the method for the generic
function as.character() must be called as.character.threshold().
The methods should have the same inputs as the generic function;
checking by calling as.character (without brackets):
## function (x, ...) .Primitive("as.character")as.character() expects two inputs: x (main object) and ... (further arguments;
unused in our case). Our as.character.threshold() should return "+" if a value
is above the limit, and "-" else.
as.character.threshold <- function(x, ...) {
res <- ifelse(x > attr(x, "limit"), "+", "-")
return(res)
}
as.character(pm25) # Testing## [1] "-" "-" "+" "+" "-"Perfect. Next, we will write the method for print() which re-uses the
as.character() method. The first input is x (see generic function print),
... will be unused again.
print.threshold <- function(x, ...) {
res <- format(x, justify = "right") # Proper formatting
res <- paste0(res, as.character(x)) # Combine 'res' with as.character(x)!!
print(res, quote = FALSE) # The print we'll see
invisible(x) # Invisible return of the original object
}
print(pm25)## [1] 100- 75- 230+ 220+ 50-We now have a very specific and easy-to-understand output of our new object class.
Note: This way (see below) it is easy to create objects of
the correct class, but with very weird content. Here we have a
character vector for our values (instead of numeric values), the
limit is not a single numeric value, and the print() method fails
(at least throws warnings).
## Warning in x > attr(x, "limit"): longer object length is not a multiple of
## shorter object length## [1] foo+ bar+ foo+Class constructor function
Much better: Create a class constructor function. This constructor (or initialization) function takes care that the objects are set up properly and only contain what they are allowed to do by providing sanity checking and intelligible error messages.
We have seen a variety of them: factor(), matrix(),
vector(), list(), to create objects of specific classes.
As we create an object of class threshold, let us define a class constructor function
threshold() with two arguments: x (data) and limit.
threshold <- function(x, limit) {
# Sanity check using stopifnot (short form of if (! ..) stop()).
stopifnot(is.numeric(x))
# 'limit' must be numeric, length one, and larger than 0
stopifnot(is.numeric(limit), length(limit) == 1L, limit > 0)
# Create our object
class(x) <- "threshold"
attr(x, "limit") <- limit
return(x)
}
pm25 <- threshold(c(100, 75, 230, 220, 50), limit = 150)
print(pm25)## [1] 100- 75- 230+ 220+ 50-This ensures that we get a proper threshold object and will
throw an error if we do something wrong.
Additional methods
To add more functionality, we can now add additional methods for our new object. In this example, we would like to have a custom summary method and two methods for plotting the data.
Summary: The summary method should show the numeric summary (as
for a numeric vector) and a count how often we are above the threshold
(using table() on the character representation of our object).
Note: the default arguments to summary() are object and ... (would
also work if you call it x).
summary.threshold <- function(object, ...) {
num <- summary(as.numeric(object)) # Numeric summary
tab <- table(as.character(object)) # Table of character representation
# Output _to console_:
cat("Numeric:\n")
print(num)
cat("\nTable:")
print(tab)
# Invisible return of both
invisible(list("numeric" = num, "table" = tab))
}
summary(pm25)## Numeric:
## Min. 1st Qu. Median Mean 3rd Qu. Max.
## 50 75 100 135 220 230
##
## Table:
## + -
## 2 3Plotting functions: To be able to graphically display the data
we would like to have two additional methods plot() and hist().
plot() should be a line plot, hist() shows us a histogram with the
distribution of the data. We specify some default arguments used to
draw the plots and forward ... to the default plotting functions
(plot() and hist()) internally. This allows us to change the
look if needed.
plot.threshold <- function(x, pch = NULL, type = "o", ...) {
# Use character representation if pch = NULL
# to plot a + or - later on.
if (is.null(pch)) pch <- as.character(x)
# Calling default plot function for numeric vectors
plot(as.numeric(x), pch = pch, type = type, ...)
# Adding a horizontal line for the limit
abline(h = attr(x, "limit"), col = "gray", lty = 2)
# Invisible return
invisible(x)
}
par(mfrow = c(1, 2)) # Subplot definition
plot(pm25) # Default
plot(pm25, col = 2, main = "Test", cex = 3) # Custom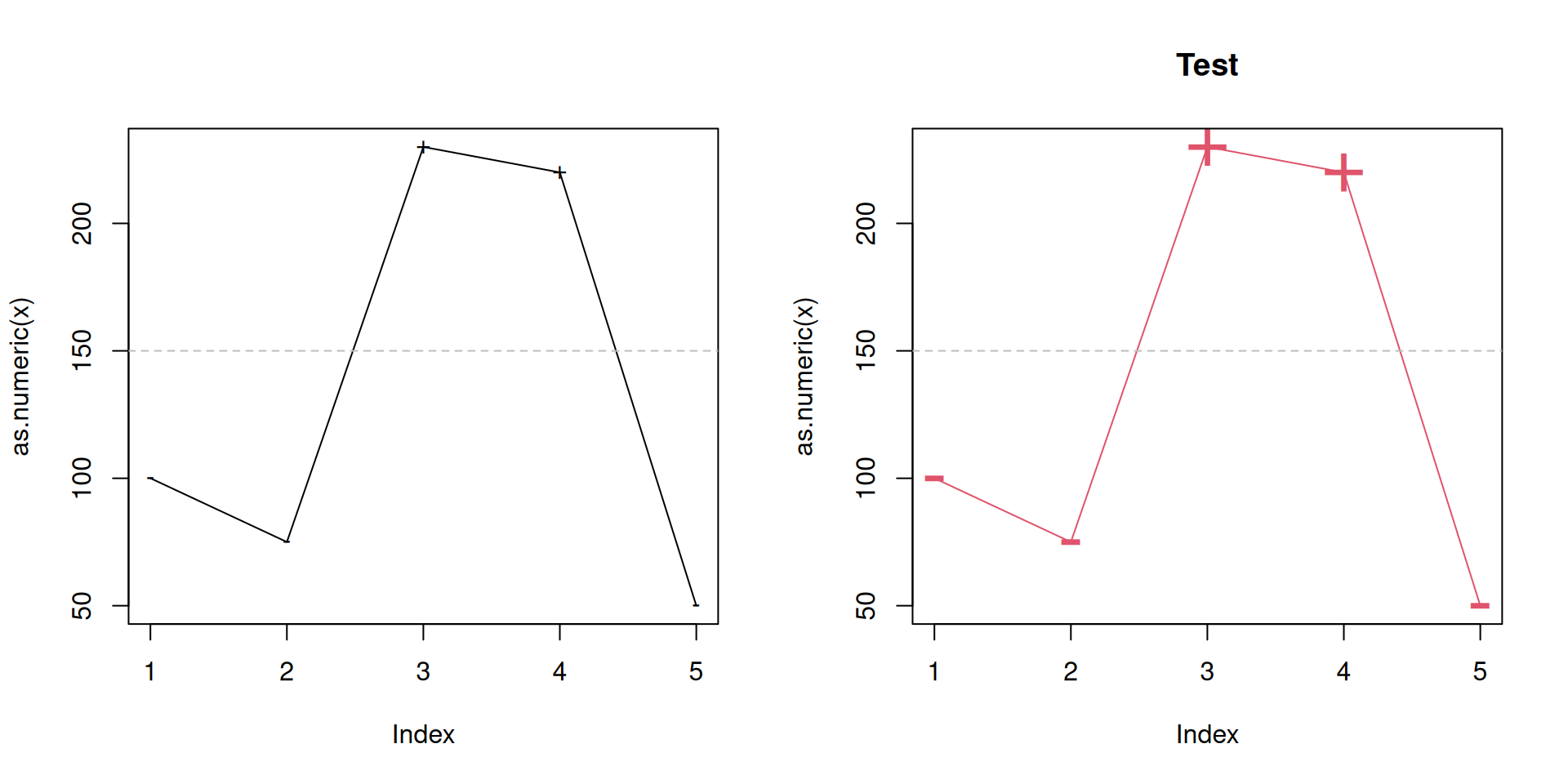
Last but not least we create a similar method for the histogram.
hist.threshold <- function(x, lcol = 2, ...) {
res <- hist(as.numeric(x), ...) # Default histogram
abline(v = attr(x, "limit"), col = lcol, lwd = 2) # Adding limit
invisible(res) # Return
}
par(mfrow = c(1, 2)) # Subplot definition
hist(pm25) # Default
hist(pm25, lcol = 3, main = "Test") # Custom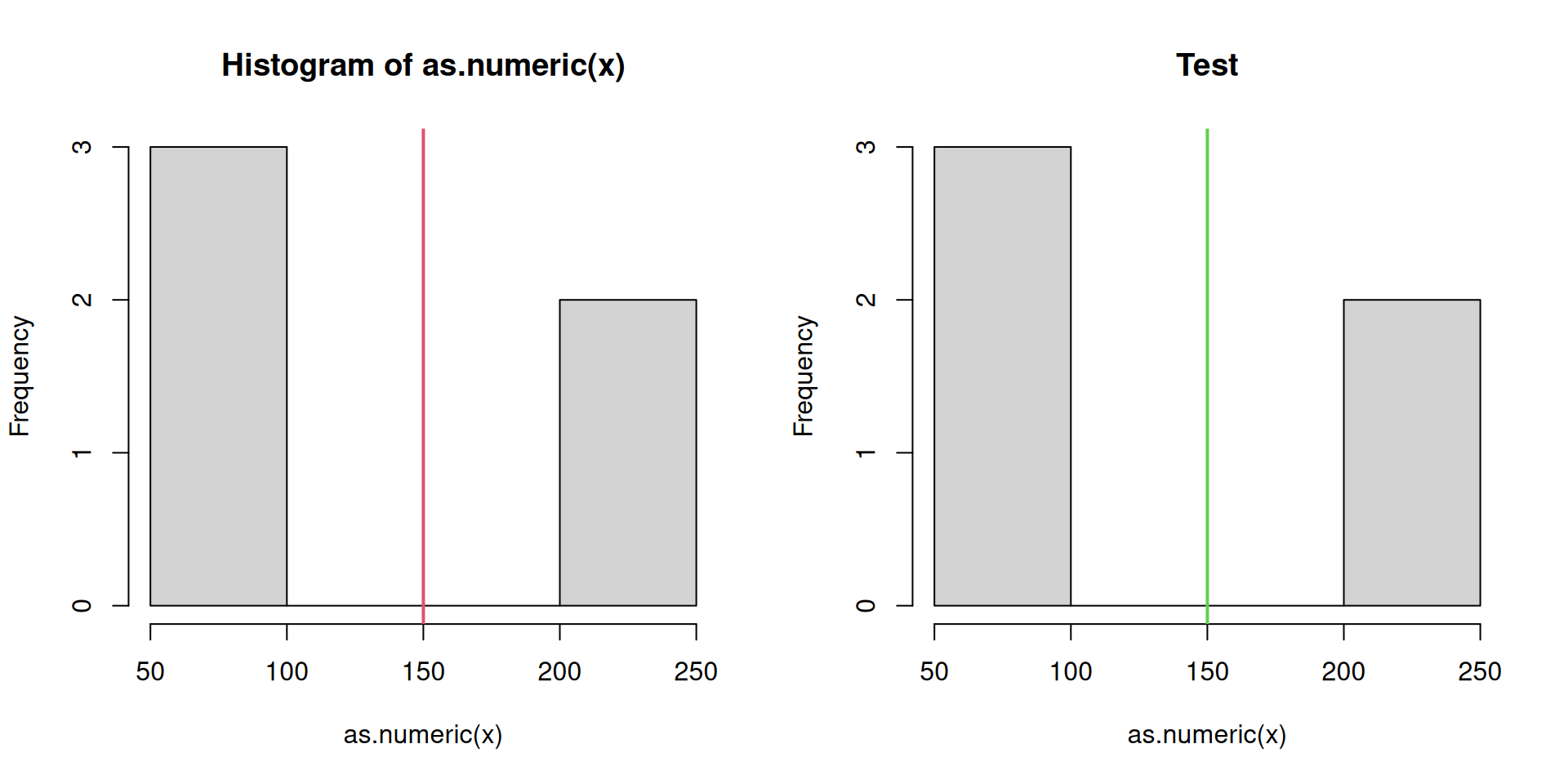
Beijing air quality
Let us now use our new object class for the data set mentioned at the beginning. The data set called “pollution-Beijing.rda” contains air pollution observations for Beijing China from 2015. To be more specific, it contains the daily average PM25 concentration. For short: the lower this value is, the lower the pollution (the better the air quality).
load("data/pollution-Beijing.rda")
plot(pollution$PM25, type = "l", col = 2, main = "PM2.5 Beijing, China, 2015")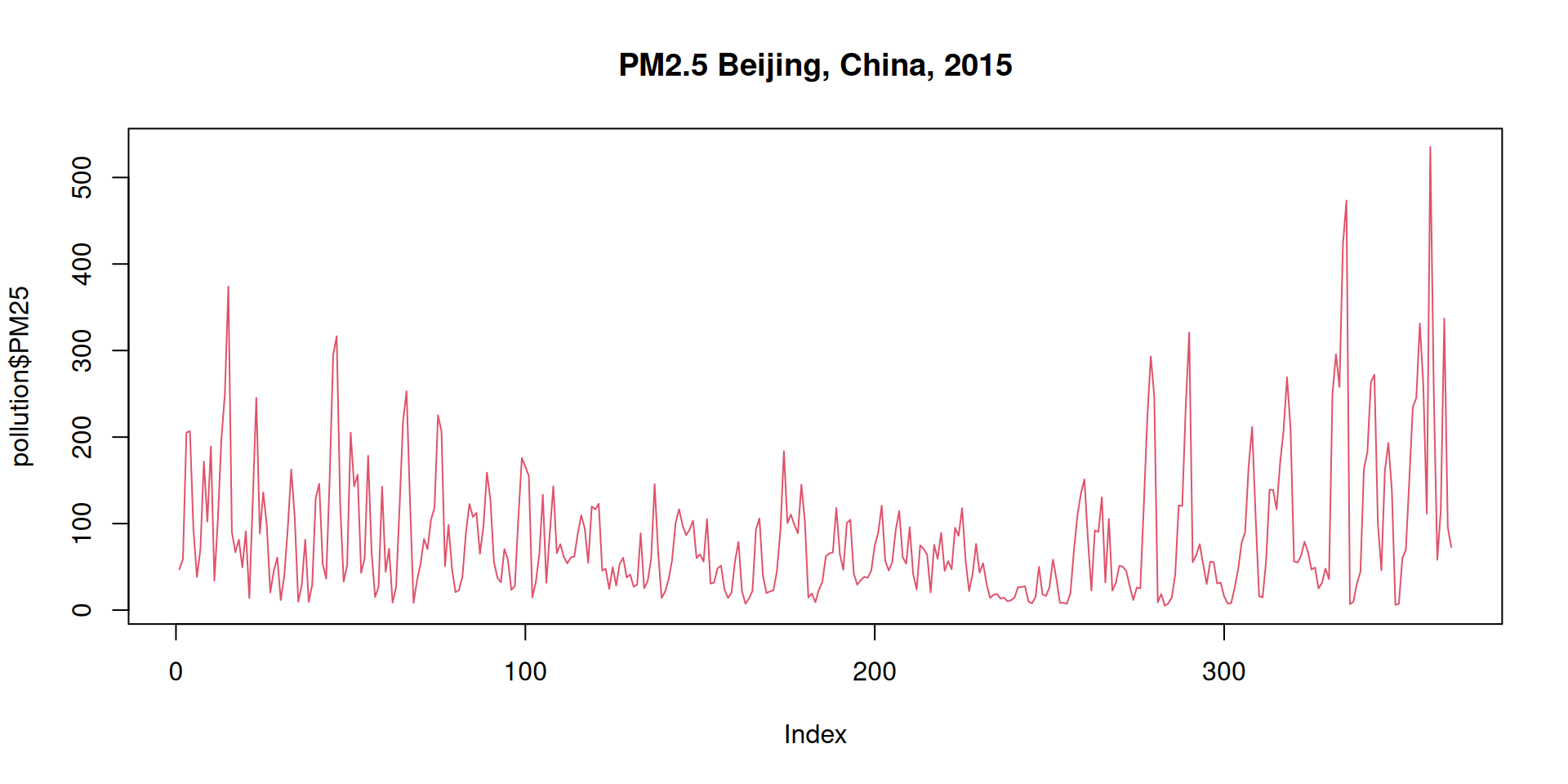
Figure 11.4: Daily mean PM25 concentration in Beijing over entire 2015.
PM25 is the concentration of particles with a diameter of about \(2.5\) micrometer, given in micrograms per cubic meter air (particular matter concentration, Feinstaubkonzentration; unit \(\mu m m^{-3}\)). It is known that these particles can cause serious health issues. Thus, the goal of the European Union was to reduce the annual (yearly) average to less than \(25\,\mu m m^{-3}\) in 2015 (\(20\,\mu m m^{-3}\) for 2020).
Let us assume that we define a limit of \(100\) as the highest allowed concentration and use our custom threshold class to process the data.
- Convert
pollution$PM25into an object of classthreshold; uselimit = 100. - Calculate the summary statistics.
- Visualize the data.
load("data/pollution-Beijing.rda") # Load data
pm25 <- threshold(pollution$PM25, limit = 100) # Create threshold object
summary(pm25) # Summary## Numeric:
## Min. 1st Qu. Median Mean 3rd Qu. Max.
## 5.10 31.50 60.50 85.86 110.40 535.30
##
## Table:
## + -
## 110 255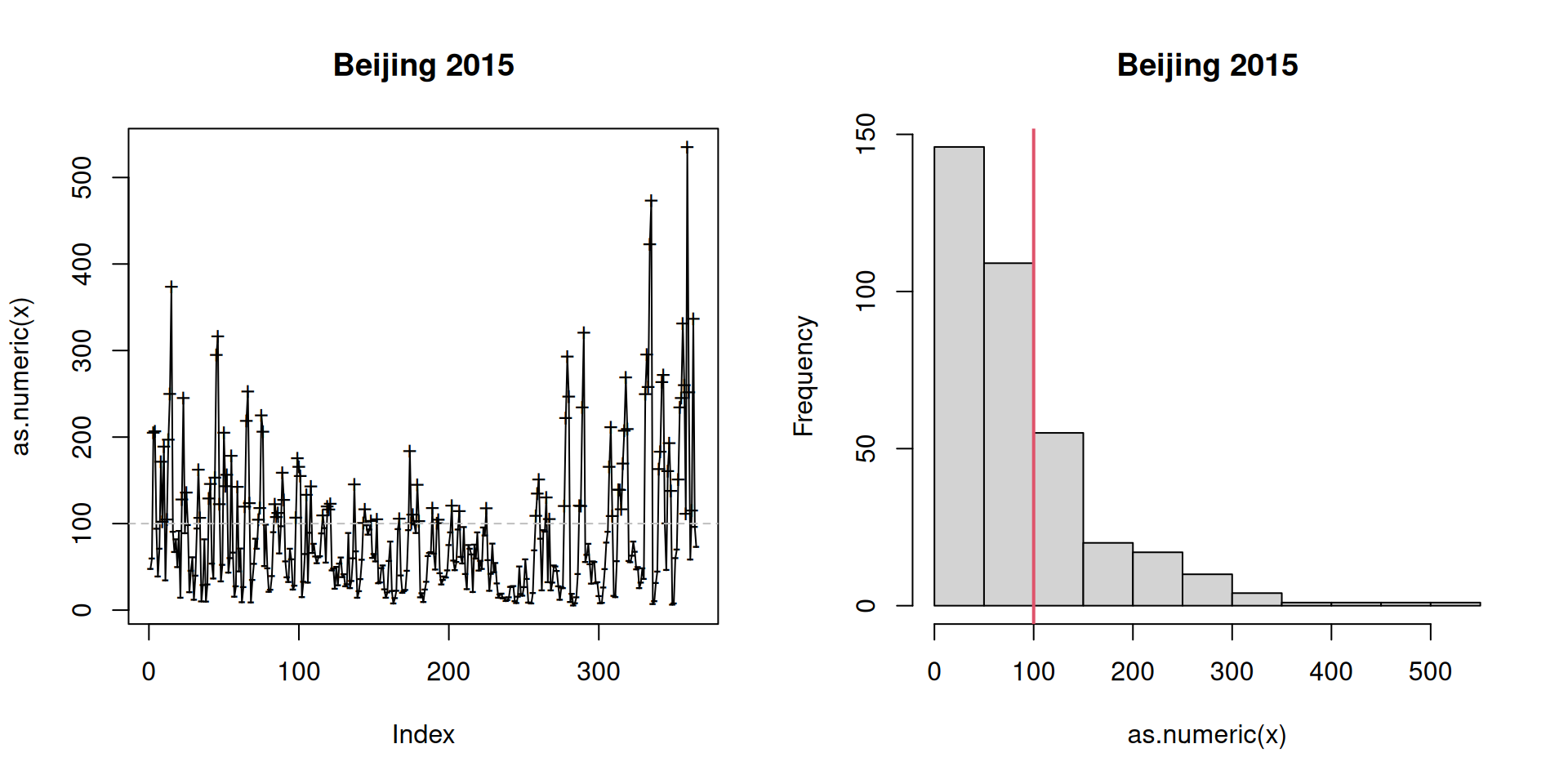
This can be very handy if there is no existing class which can handle your data or you need a more efficient way to handle your own data.
Exercise 11.1 Letters class: As another example to practice, let us create a
simple letter class. A class for a single letter (a-z, A-Z).
In this case, we would like to have a new class letter which consists
of one single character value.
Class constructor function
Input must be a single character which is in letters (a-z) or LETTERS (A-Z).
All other inputs will result in an error.
letter <- function(x) {
# Sanity checks
if (!is.character(x)) stop("Input 'x' must be of class character.")
if (!length(x) == 1L) stop("Input 'x' must be of length 1")
if (!x %in% letters & !x %in% LETTERS) stop("'x' is no valid letter")
# Now append class and return the new object
class(x) <- "letter"
return(x)
}
# Testing
(x <- letter("A"))## [1] "A"
## attr(,"class")
## [1] "letter"## Error in letter(1L): Input 'x' must be of class character.## Error in letter("1"): 'x' is no valid letterCustom print() method
print.letter <- function(x, ...) {
cat("This is a letter:", dQuote(x), "\n")
invisible(x)
}
print(x)## This is a letter: "A"Summary and structure
In this case, we are happy how the default methods str() and summary()
look like, thus we don’t really need to create a custom one. If needed,
we could of course write our own str.letter() and summary.letter() methods.
## 'letter' chr "A"## Length Class Mode
## 1 letter character11.4 Date and date time
We now better understand how different S3 objects are constructed and
processed. Let us have a closer look at some other very useful objects:
Date and POSIXt. These objects represent dates and date-time and
allow for arithmetics (e.g., how many days until new year?).
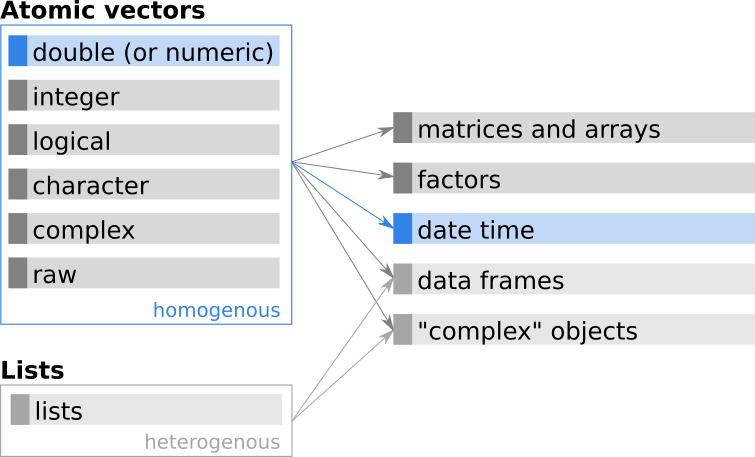
Figure 11.5: Additional base R classes based on atomic vectors.
Technically, a Date or POSIXt object is nothing else than a
simple numeric vector, but displayed as a human-readable date (like a string),
similar to the factor class.
For both classes a wide range of methods exist which makes it very easy to work with. We’ll only cover few in this chapter. Run the following two commands yourself to see what’s possible.
Dates
- Calendar dates: Are stored as objects of class
Date. No time information. - Structure: Numeric vector; the number of days since
1970-01-01plus a class attribute. - Creation: Typically from strings (ISO 8301 format; same is used for printing).
## [1] "2021-01-01"## [1] "2021-03-29"## class typeof
## "Date" "double"As shown, d is an object of class "Date", type "double", in this case of length 1.
If we call unclass(d) all classes are removed which allows us to ‘look inside’ the
structure of the object.
## [1] 18628As shown, this is all. It’s nothing else than the numeric value
\(18628\) and
a class attribute "Date" (which we just removed here).
Working with dates
The nice thing is that we easily work with this object and e.g., add or subtract integers. E.g.:
- Day before
d:d - 1. - Date in 7 days from now:
d + 7.
The same also works with integer sequences, e.g., seven days starting with d (2021-01-01):
## [1] "2021-01-01" "2021-01-02" "2021-01-03" "2021-01-04" "2021-01-05"
## [6] "2021-01-06" "2021-01-07"Alternatively we can make use of the seq() function to e.g.,
create a decreasing sequence with a seven-day interval:
## [1] "2021-01-01" "2020-12-25" "2020-12-18" "2020-12-11" "2020-12-04"Multiple date objects can also be used to do some calculations. If we are interested in the number of days between two dates, we simply subtract one from another.
## Time difference of 87 days## [1] 87Formatting dates
To format objects of class Date (e.g., for pretty printing or additional processing
steps) we can use the generic function format(). This generic is not solely used for
for objects of class Date, but many other classes (try methods(generic.function = "format")).
This chapter will only cover the use of the generic function format() for objects of class
Date. Let us define a new object birthday as follows:
## [1] "2017-02-01"## [1] "Date"format() can now be applied to convert the object into a representation of choice.
The function always returns a character vector, wherefore you will lose some
functionality (e.g., arithmetics is no longer possible on character representations
of a date). However, it can be used for pretty printing or bring your date information
into a different format.
When called without any additional arguments, format() will return the date information
as an ISO representation. While it looks similar to the original object, it is now
of class character.
## [1] "2017-02-01"## [1] "character"Custom formatting can be defined by providing the additional format
argument. The code below, for instance, returns the name of the month or the
name of the day from the object birthday:
## [1] "February"## [1] "Wednesday"## [1] "2017-02-01"## [1] "01.02.2017"## [1] "His/her birthday is on Wednesday, February 01."The argument format expects a character which can contain simple text (see
above) and/or some formatters like %Y, %m, %d, %B, %A. A few
examples are listed in the table below, but many more are available. For a
full list of possible formats have a look at the details section of
?strftime.
Note that some of them (e.g., "%B",“%A”`) depend on your system language
(not necessarily in English).
| Argument | Description | Result |
|---|---|---|
"%a" |
Abbreviated weekday | "Wed" |
"%A" |
Full weekday | "Wednesday" |
"%b or "%h" |
Abbreviated month | "Feb" |
"%B" |
Full month | "February" |
"%d" |
Day of the month | "01" |
"%j" |
Day of the year | "032" |
"%m" |
Month | "02" |
"%U" |
Week | "05" |
"%x" |
Date | "02/01/2017" |
"%Y" |
Year with century | "2017" |
Time
Note that objects of class Date do not contain any time information.
To handle date and time we need objects of class POSIXt. They work
the very same as Date objects expect that an increment of +1 is
no longer +1 day, but +1 second. This is flagged as additional
(optional) content but works the very same as the Date class (including
arithmetics or formatting via format()).
Thus, if you are interested in this topic, feel free to read the following ‘excursion’.
- Date/time: Stored as objects of class
"POSIXct"(inheriting from"POSIXt"); date and time. - Time zone: As dealing with time, the time zone is getting important. If not set, the computer default time zone is used.
- Structure: Numeric (double) vector, number of days since 1970-01-01 00:00:00 UTC (“UNIX epoch”), a class attribute, and an additional time zone attribute.
- Creation: Typically from strings, defaulting to ISO 8301, also used for printing.
Important to keep in mind: the time zone of your computer is used. The standard time
zone of our system (where the book is compiled) uses "UTC" (universal time code).
We can explicitly define the time zone using tz = "..." if needed.
## [1] "2021-01-01 UTC"## [1] "2021-01-01 CET"Checking properties/attributes of dt1:
## $class
## [1] "POSIXct" "POSIXt"
##
## $typeof
## [1] "double"
##
## $attributes
## $attributes$class
## [1] "POSIXct" "POSIXt"
##
## $attributes$tzone
## [1] ""As POSIXct inherits from POSIXt we can see that there are two classes
(like for matrices c("matrix", "array")).
Let us unclass both objects to see how they are constructed:
## [1] 1609459200
## attr(,"tzone")
## [1] ""## [1] 1609455600
## attr(,"tzone")
## [1] "CET"As shown, both objects are numeric (double) vectors but contain different numeric values. The numbers always represent the number of seconds to the reference date 1970-01-01 00:00:00 UTC (UTC time zone). As there is one hour offset between “UTC” and “CET”, there is a one-hour difference between these two objects.
## Time difference of 1 hoursAs for converting an object of class character to Date, as.POSIXct accepts
a custom format definition in case our date and time information (the input)
does not come in a standard ISO represesentation.
## [1] "2020-03-29 12:47:04 UTC"Working with date/time
As with Date objects we can now work with the POSIXct object(s).
Note that, internally, they count the number of seconds. Thus, adding +1
will increase the time by 1 second (not one day as for Date objects).
## [1] "2020-12-31 23:59:00 UTC"## [1] "2021-01-01 03:00:00 UTC"## [1] TRUE## [1] FALSEFormatting POSIXt objects
Formatting objects of class POSIXt works in the same way as formatting
objects of class Date. Internally, R is now using the dedicated methods
format.POSIXct() (or format.POSIXlt(); not covered here) instead of
format.Date() as used for objects of class Date. However, the usage
is barely different:
## [1] "2021-01-01"## [1] "2021-01-01 00:00:00"## [1] "Friday 01, January 2021"## [1] "Object `dt1` (2021-01-01 00:00:00 UTC) belongs to Julian day 001"## [1] "1 hours"Or to get a bit more creative (if needed):
## [1] "00h01m (on Friday January 01, 2021)"Plotting
Date and date-time objects are very handy when working with date/time-related data. As an example, let us imagine we have some (random) observations over one year and would like to plot this.
# Generate Date vector over entire 2021
dates <- seq(from = as.Date("2021-01-01"),
to = as.Date("2022-01-01") - 1,
by = 1L)
c(min(dates), max(dates))## [1] "2021-01-01" "2021-12-31"# Create some random data -> add both to a data frame.
set.seed(6020) # Pseudo-randomization
df <- data.frame(date = dates, value = rnorm(length(dates)))
head(df)## date value
## 1 2021-01-01 -0.3421647
## 2 2021-01-02 0.4528670
## 3 2021-01-03 0.6169125
## 4 2021-01-04 -0.6121245
## 5 2021-01-05 -1.2603337
## 6 2021-01-06 -0.4266803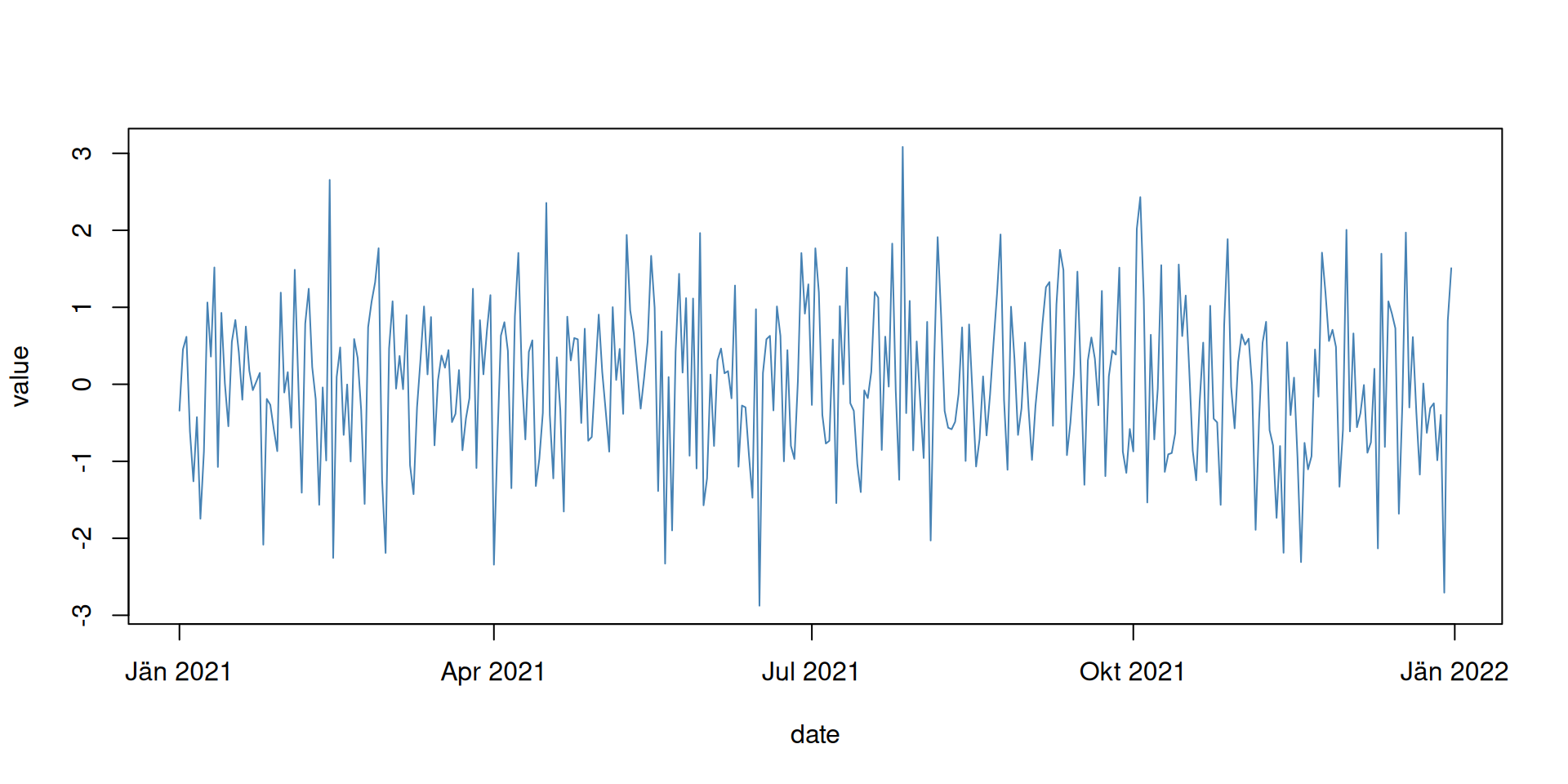
Or slightly different using type = "h" with different colors for
negative/positive values:
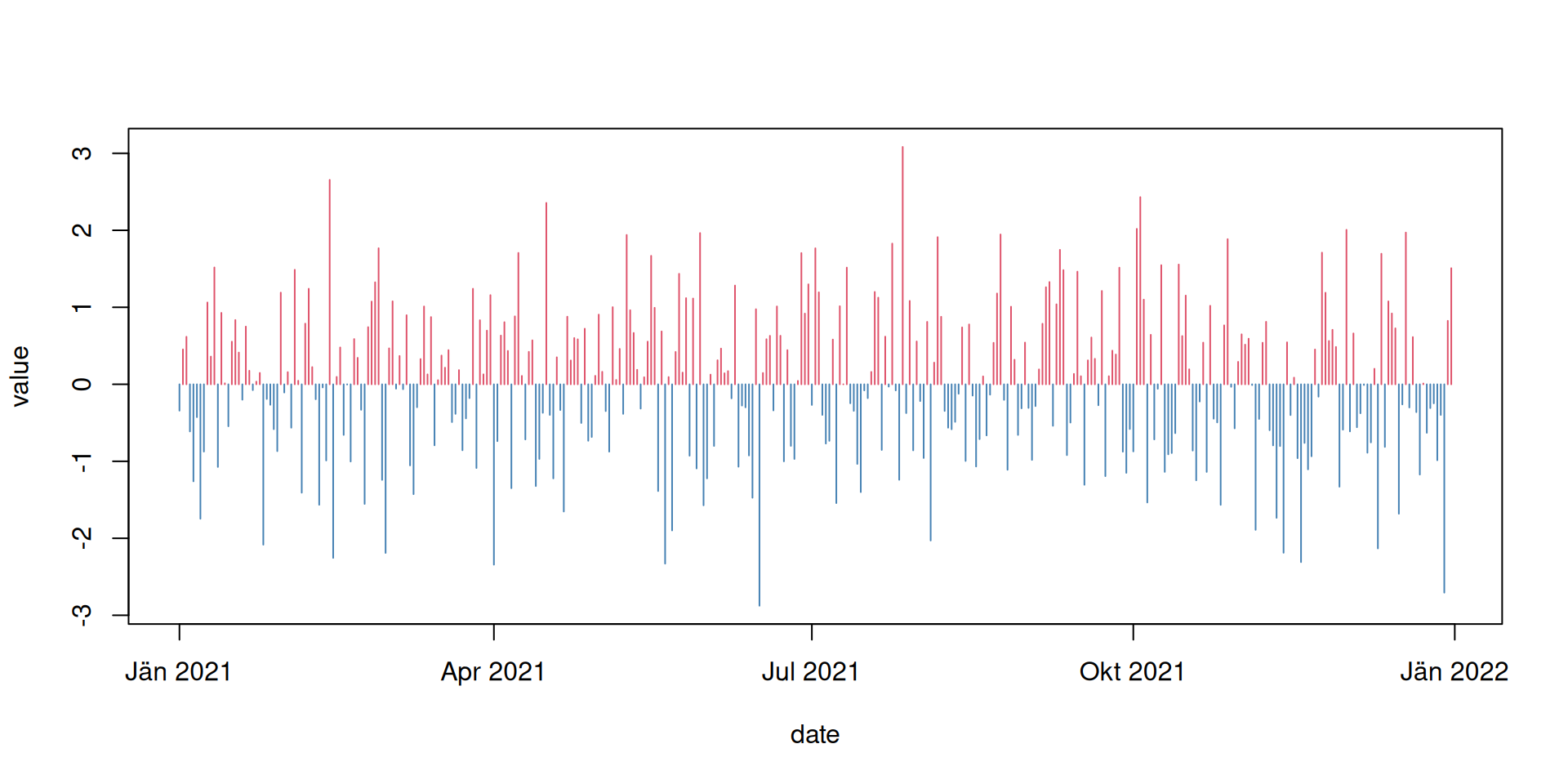
As you can see, R now takes care to plot the data correctly along the x-axis and also adds useful labels. In addition, date and date-time objects are super useful to e.g., calculate annual (yearly), monthly, or daily values (e.g., mean, minimum, maximum) and much more.